According to the World Health Organisation (WHO), over 1500 new pathogens have been found since the 1970s, and almost 40 new infectious diseases have been diagnosed. Many diseases have had a significant impact on populations, with numerous major outbreaks happening in the last 20 years, including SARS (2002–2003), Ebola (2014–2016), Swine flu (2009–2010), Zika virus (2015–2016), and more recently COVID-19 (2019–2021). Drug-resistant microorganisms have also been connected to the re-emergence of infectious illnesses.

Early detection of disease outbreaks at the community level plays a key role in controlling the spread of the disease and can prompt intervention for the reduction of morbidity and mortality that may result from it, as well as the application of preventive and control measures. Public health requires novel monitoring and management approaches for the prevention of infectious diseases within a community. These new technologies should be flexible, cost-effective, and scalable and should provide comprehensive and objective data in real time.
Wastewater-based epidemiology (WBE), which can provide objective and comprehensive real-time assessments of public and environmental health status, has gained a large interest during the COVID-19 pandemic. WBE is a new epidemiology tool to measure the spread of diseases, drug use and other biological or chemical indicators in a population. WBE has the potential to act as a complementary approach for current infectious disease surveillance systems and can inform and improve public health responses through the analysis of population pooled wastewater samples.
Several methods and workflows are currently in use for monitoring biomarkers in wastewater. Each method may vary slightly but most of them consist of four main steps as shown in the workflow below, in which (1) a sample is collected at near-source or at wastewater treatment plant and (2) transferred to a testing laboratory for analysis. Once received by the laboratory, (3) the samples are tested for the presence of the target biomarkers and (4) the obtained results are analysed and combined with other information to interpret the data.
-
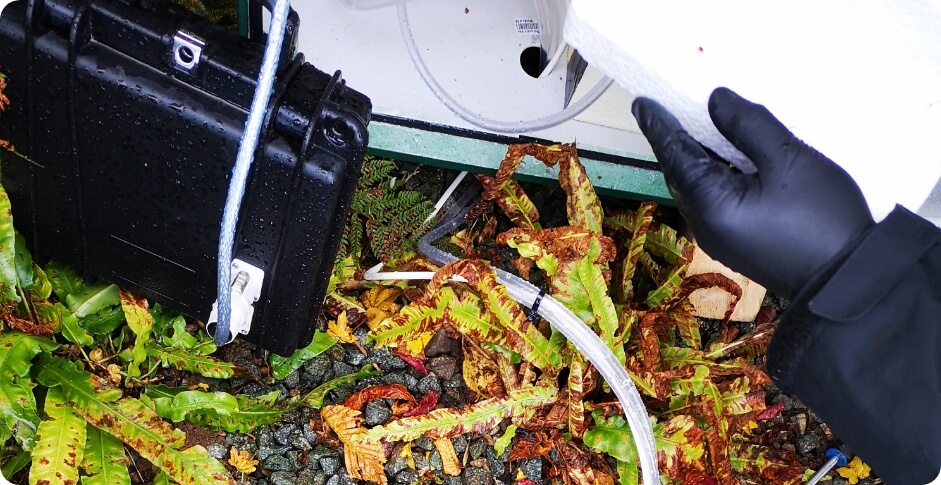
Stage 1 : Wastewater Sampling
-
Stage 2 : Samples are transported to the labratory
-
Stage 3 : Sample analysis
-
Stage 4 : Data Interpretation
Two types of wastewater samples are suitable for WBE:
- untreated wastewater, which includes waste from household or building use and contains human faecal waste
- primary sludge, which consists of the suspended solids that settle out of wastewater during the first solids removal (called “sedimentation”) within the wastewater treatment facility.
Samples are generally collected following either the grab, the composite or the passive methods depending on the constraints associated with the collection point (e.g. accessibility, access to electricity), as well as the goal of the surveillance program. Grab samples are directly taken from a sewer manhole or other location within the sewer shed, often by dipping a beaker or bottle into the sewage to collect the sample. Although rapid and simple, this method only offers a snapshot of the conditions at the time of collection and is highly influenced by daily fluctuations in wastewater flow and composition. To circumvent this problem, composite samples are collected by pooling multiple grab samples at a specified frequency over a set period. They represent the average wastewater characteristics during the period of collection and are used in most WBE studies. Finally, passive sampling uses a physical device that contains absorbent materials or membranes placed in a targeted sewage catchment to capture viruses and other particles for a defined period. Passive samplers are easy to deploy and collect from small catchments and do not require power to operate, and thus can be used in any accessible sewage line.
20/30 Labs play an important role in the effort to establish WBE as an important part of a national surveillance method. During the Covid-19 pandemic, we worked on many different projects and ran pilot studies, monitoring Care Houses, airports and residential areas. We ran thousands of tests and tested all the different sampling methods, to looking for the presence of SARS-CoV-2 in the sample. We also tested for Influenza A & B, RSV, Norovirus G1 & G2, Ammonium, Orthophosphate and pH, further establishing the approach as efficient, rapid and simple.
REFERENCES:
- Baker, R.E., Mahmud, A.S., Miller, I.F. et al. - Infectious disease in an era of global change. Nat Rev Microbiol 20, 193–205 (2022). https://doi.org/10.1038/s41579-021-00639-z
- Steele L, Orefuwa E, Bino S, Singer SR, Lutwama J, Dickmann P. Earlier Outbreak Detection - A Generic Model and Novel Methodology to Guide Earlier Detection Supported by Data From Low- and Mid-Income Countries. Front Public Health. (2020) Sep 11;8:452. doi: 10.3389/fpubh.2020.00452.
- Adhikari, S., Halden, R.U. - Opportunities and limits of wastewater-based epidemiology for tracking global health and attainment of UN sustainable development goals. Environment International, Vol.163, (2022). https://doi.org/10.1016/j.envint.2022.107217.