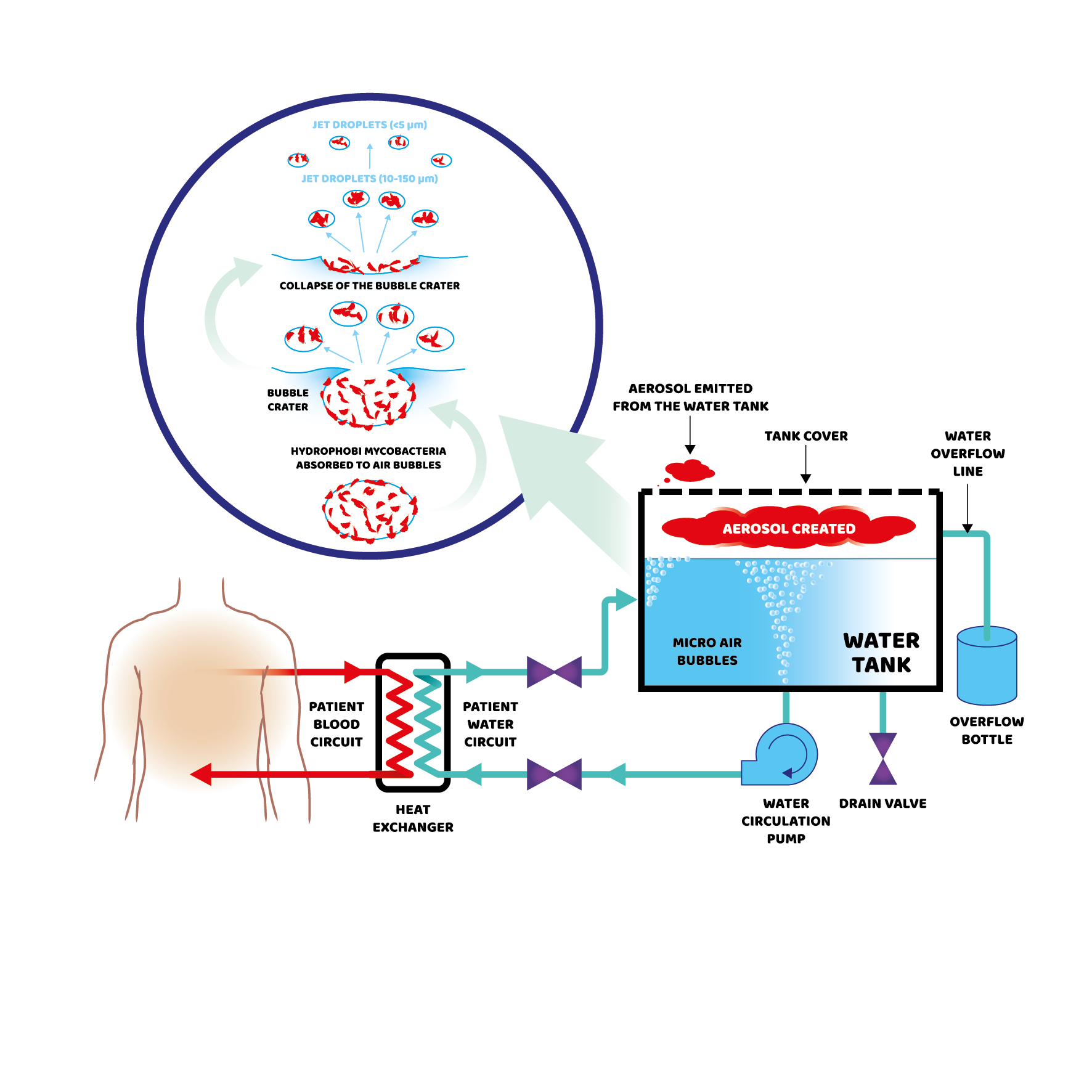
A global medical solutions company, that supplies bathing systems into the NHS, had a contamination outbreak at several UK sites. During routine water testing by 20/30 labs, Pseudomonas aeruginosa was isolated from the faucets and shower heads, but not the supply water. The baths were installed and tested prior to patient contact and without contamination in the incoming water, this suggested colonisation at the manufacturing site (in Europe). Water was sampled from the European site and tested, and the detected contamination was independently determined as caused by Pseudomonas aeruginosa ST-17 using WGS followed by Multi-Locus Sequence Typing (MLST).

The Pseudomonas aeruginosa colonies isolated at 20/30 labs were re-plated for pure culture and confirmed using MALDI-TOF bacterial identification. Whole genome sequencing (WGS) was used to determine the relationship between these colonies and the strain at the manufacturing site. Extraction of genomic DNA (gDNA) and NGS library preparation was carried out. Post base-calling, Multi-Locus Sequence Typing (MLST) was conducted, for 7 housekeeping genes. This MLST method was used at the request of the client, to remain consistent with their previous approach, and 20/30 Labs was able to facilitate this. Consensus was found between the isolates, with the UK isolate also determined to be Pseudomonas aeruginosa ST-17, confirming a source outbreak. This information was used by the medical solutions company to determine whether to deep clean and relocate the manufacturing site.
20/30 Labs can provide very quick turnaround WGS, and our reports provide a detailed explanation of what this means to the client. By approaching genomic and molecular studies through the lens of a microbiologist we are expertly placed to not only provide the highly technical expertise required for WGS but also go through what the result means for you and your contamination event.
Approach
-
Stage 1 : Objective
The client supplied a water sample(s) for the purposes of WGS.
20/30 Labs was to use the Nanopore GridION platform followed by Multi-Locus Sequence Typing (MLST) on the consensus genome produced, to confirm isolated organism is Pseudomonas aeruginosa ST-17 i.e., colonisation from the manufacturing site.
-
Stage 2 : Colony identification and isolation
Following membrane filtration and incubation, samples exhibited growth as expected, representative colonies were picked and analysed via the Bruker Microflex LT/SH MALDI-TOF platform. The MALDI-TOF system is the optimal solution for fast and accurate bacterial identification, and 20/30 labs hold an extensive customised library.
Subsequently, single characteristic colonies were isolated from the sample plate and streaked for single colonies of pure culture, which were then used for extraction of genomic DNA (gDNA).
-
Stage 3 : gDNA Extraction
Extraction was performed using the Qiagen Genomic-tip system, as per Nanopore (manufacturers) IFU. Throughout, no fragmentation of the gDNA was undertaken.
-
Stage 4 : NGS Library Preparation and Sequencing.
Following extraction, NGS library preparation was performed. The extracted gDNA was sequenced directly and without prior PCR amplification. To do so, the extracted gDNA was first normalised, end-repaired, and then end-prepped for ligation of the Nanopore adaptor molecules. 10 femtomoles of the prepared library was then immediately loaded onto a nanopore flow cell and sequencing initiated with the specific run parameters. The sequencing application resulted in the generation of 15.86 Gb of read data with 1.12million unique reads. Following de novo genome assembly using the Flye assembler tool, mean sequencing depth was determined to be approximately 1800X.
-
Stage 5 : Multi-Locus Sequence Typing
Post base-calling, Multi-Locus Sequence Typing (MLST) was conducted for the sample:
Reference alleles for the seven Pseudomonas aeruginosa MLST genes provided by the client (trpE, nuoD, aroE, acsA, guaA, ppsA, mutL) were downloaded from pubmlst.org. The tool krocus was then used to map sample reads to the reference alleles. The MLST type of the isolate was determined by comparing the present alleles with the MLST profiles list for Pseudomonas aeruginosa, downloaded from pubmlst.org. -
Stage 6 : Result
Consensus was found between the isolates, with the UK isolate also determined to be Pseudomonas aeruginosa ST-17, confirming a source outbreak.